Overview
All cells decode information from their surroundings via receptor molecules, which are proteins that detect and translate these signals to the language of the cell, thereby altering its function. Sensory nerves are specialists of this process; they monitor both external and internal environments and relay this information to alter the behavior of an entire organism. They come in many different flavors, expressing different combinations of receptor molecules, which in turn determine their response profile. Many of the genes that encode sensory receptors are still unknown, their potential as therapeutic targets as yet untapped. We are interested in identifying and characterizing these molecules, to understand their role in health and diseases such as chronic pain.
Experimental approaches
Current projects
We study ion channels and GPCR-s that decode noxious physical and chemical cues from the environment in primary sensory neurons using a combination of electrophysiology, fluorescent imaging and molecular biology. Our aim is to identify target proteins and signaling pathways that allow intercepting painful stimuli at the level of the peripheral sensory nerve.
More
Receptor molecules that enable primary sensory neurons to respond to noxious stimuli are prime targets for next-generation analgesic therapy. Intercepting the pain signal at this level promises both relative safety and efficacy. The prevalence of chronic pain conditions, and the addictive potential of opioids make this research goal a high priority. We use patch clamp electrophysiology combined with fluorescent imaging techniques and molecular biology approaches to study the regulation of receptors and ion channels that are crucial for the detection of sensory stimuli in primary sensory neurons of Dorsal Root Ganglia (DRG). We are interested in receptors of extreme temperature and mechanical stimuli, as well as natural compounds that elicit complex sensory phenomena.
Targeted cellular evolution

We develop novel approaches to enable high-throughput screening via genome-wide mutagenesis. We use high-yield screening tools to identify new molecules and pathways in the sensory nervous system, as well as to conduct large-scale drug re-purposing screens to identify off-target effects of both previously characterized molecules and novel chemical compounds and fragments.
More
 Coupling cellular function to its genetic underpinning is a challenging task. This project, funded by HORIZON 2020, BBSRC, and Wellcome Trust, aims to design experiments that harness the logic of evolution to achieve this goal. We use genome editing tools to induce random changes in the genetic makeup of cells, occasionally giving rise to rare emergent cellular phenotypes. We then conduct an unbiased selection process that allows us to enrich these rare cells and study their genetic makeup. We use this approach to uncover the identity of molecular receptors in a wide variety of biological fields.
Coupling cellular function to its genetic underpinning is a challenging task. This project, funded by HORIZON 2020, BBSRC, and Wellcome Trust, aims to design experiments that harness the logic of evolution to achieve this goal. We use genome editing tools to induce random changes in the genetic makeup of cells, occasionally giving rise to rare emergent cellular phenotypes. We then conduct an unbiased selection process that allows us to enrich these rare cells and study their genetic makeup. We use this approach to uncover the identity of molecular receptors in a wide variety of biological fields.
Past projects
Mechanosensors in proprioceptive neurons
Proprioceptive neurons signal positional cues of limbs and torso to the brain, making complex, coordinated movements possible. The mechano-sensitive ion channel Piezo2 is the principal mechanical receptor in proprioceptive neurons enabling this critical function. See Publications
More
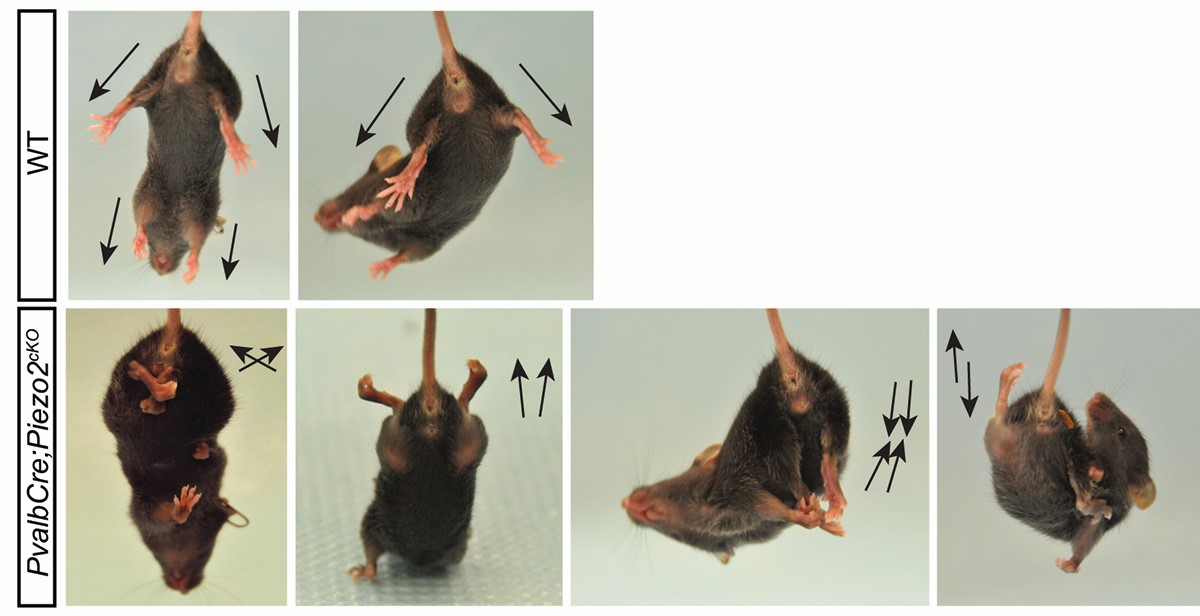 The mechanosensitive ion channel Piezo2 plays a critical role in somatosensory function. This channel allows not only the detection of innocuous touch, but is also expressed in nerves that detect the state and change of muscle and tendon stretch. The latter neurons conveys positional information to the central nervous system, which is critical for the precise execution of complex movements. With the help of these neurons we can maintain posture and execute tasks even without visual feedback. Lack of Piezo2 leads to loss of such coordination both in both mice and humans (Woo et al. 2015).
The mechanosensitive ion channel Piezo2 plays a critical role in somatosensory function. This channel allows not only the detection of innocuous touch, but is also expressed in nerves that detect the state and change of muscle and tendon stretch. The latter neurons conveys positional information to the central nervous system, which is critical for the precise execution of complex movements. With the help of these neurons we can maintain posture and execute tasks even without visual feedback. Lack of Piezo2 leads to loss of such coordination both in both mice and humans (Woo et al. 2015).
Mechanotransduction in red blood cells
Red blood cells are exposed to various kinds of mechanical agitation in the vasculature. Mechanosensitive Piezo1 proteins allow these cells to detect and adapt to these forces, enabling them to squeeze through very narrow passages. Piezo1-dependent volume regulation is also a fascinating new factor in malarial susceptibility. See Publications
More
 Red blood cells squeeze through capillaries whose diameter is often smaller than that of the RBC itself. Passage through such narrow vasculature conveys mechanical stress on the RBC, which triggers an active signaling cascade to initiate the loss of a small amount of volume, facilitating erythrocyte passage through the smallest of capillaries. This signaling is initiated by the mechanosensitive ion channel Piezo1. Calcium entry through this channel activates “Gárdos” calcium-activated potassium channels, triggering cell dehydration driven by potassium and chloride efflux from the cell. Gain of function mutations of Piezo1 lead to a clinically benign condition of mildly reduced RBC size. Interestingly, some of these variants also convey resistance to malarial infection, putting Piezo1 on the map of potential targets for treatment of this disease (Cahalan and Lukacs et al, 2015).
Red blood cells squeeze through capillaries whose diameter is often smaller than that of the RBC itself. Passage through such narrow vasculature conveys mechanical stress on the RBC, which triggers an active signaling cascade to initiate the loss of a small amount of volume, facilitating erythrocyte passage through the smallest of capillaries. This signaling is initiated by the mechanosensitive ion channel Piezo1. Calcium entry through this channel activates “Gárdos” calcium-activated potassium channels, triggering cell dehydration driven by potassium and chloride efflux from the cell. Gain of function mutations of Piezo1 lead to a clinically benign condition of mildly reduced RBC size. Interestingly, some of these variants also convey resistance to malarial infection, putting Piezo1 on the map of potential targets for treatment of this disease (Cahalan and Lukacs et al, 2015).